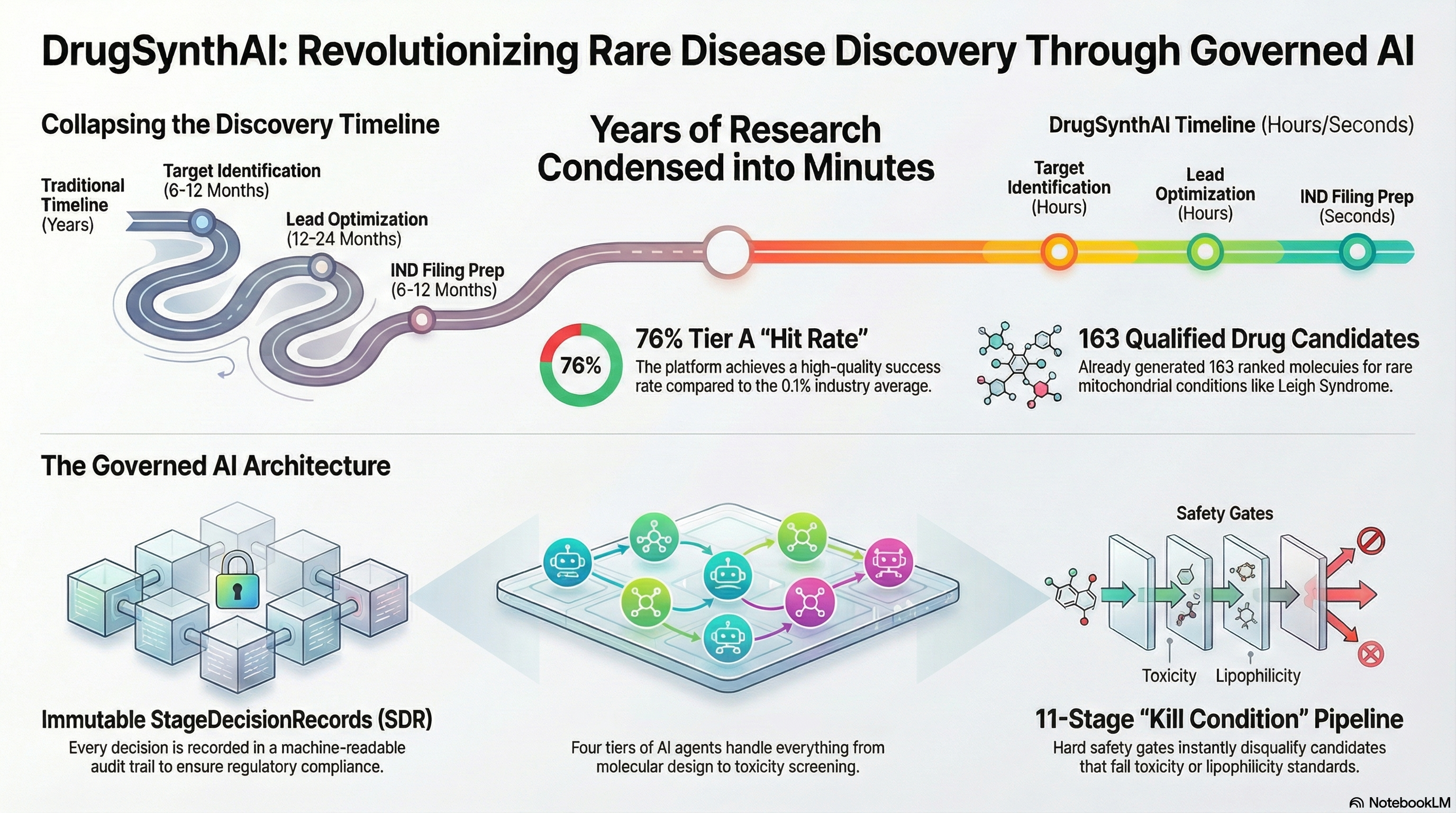
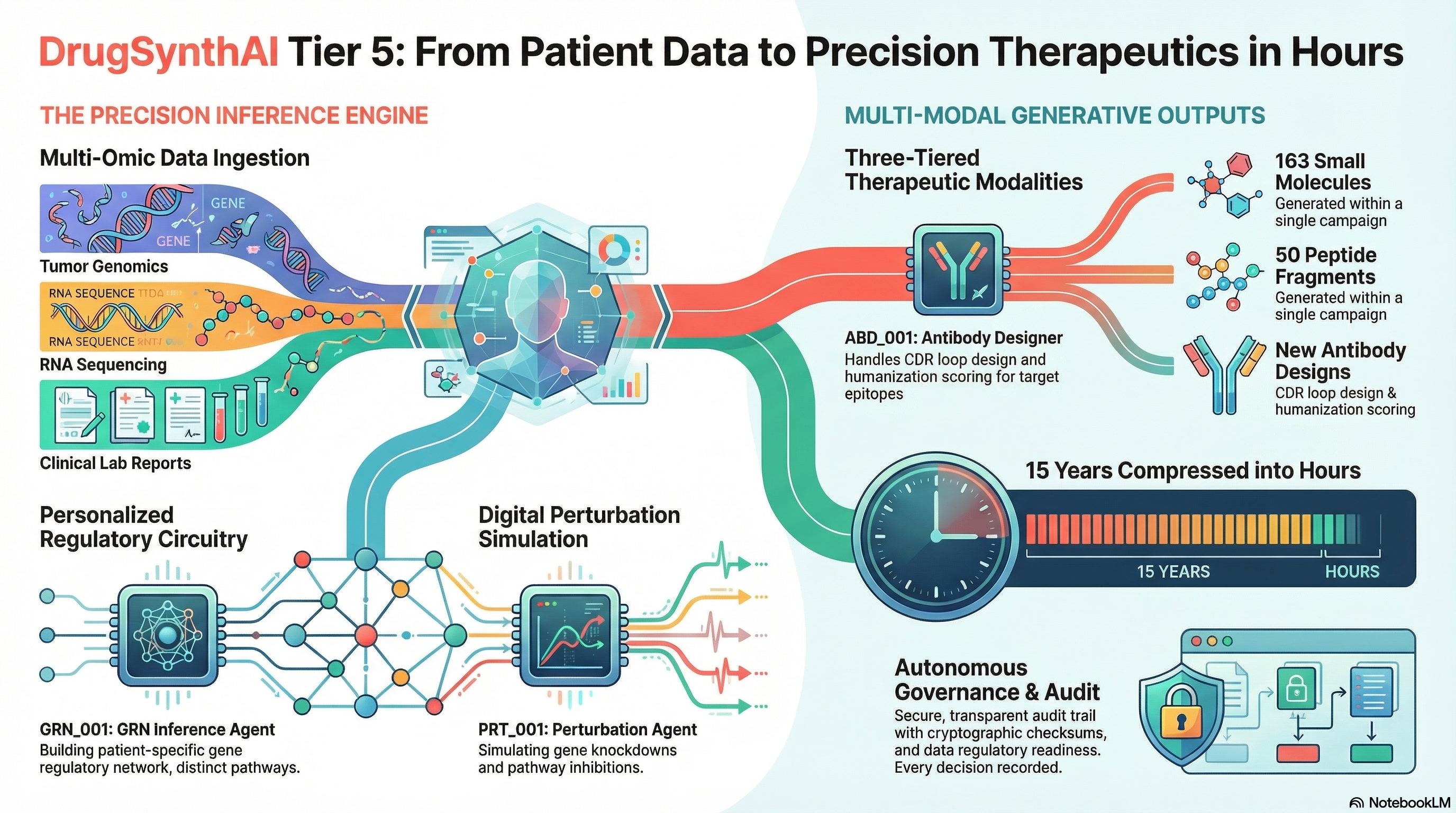
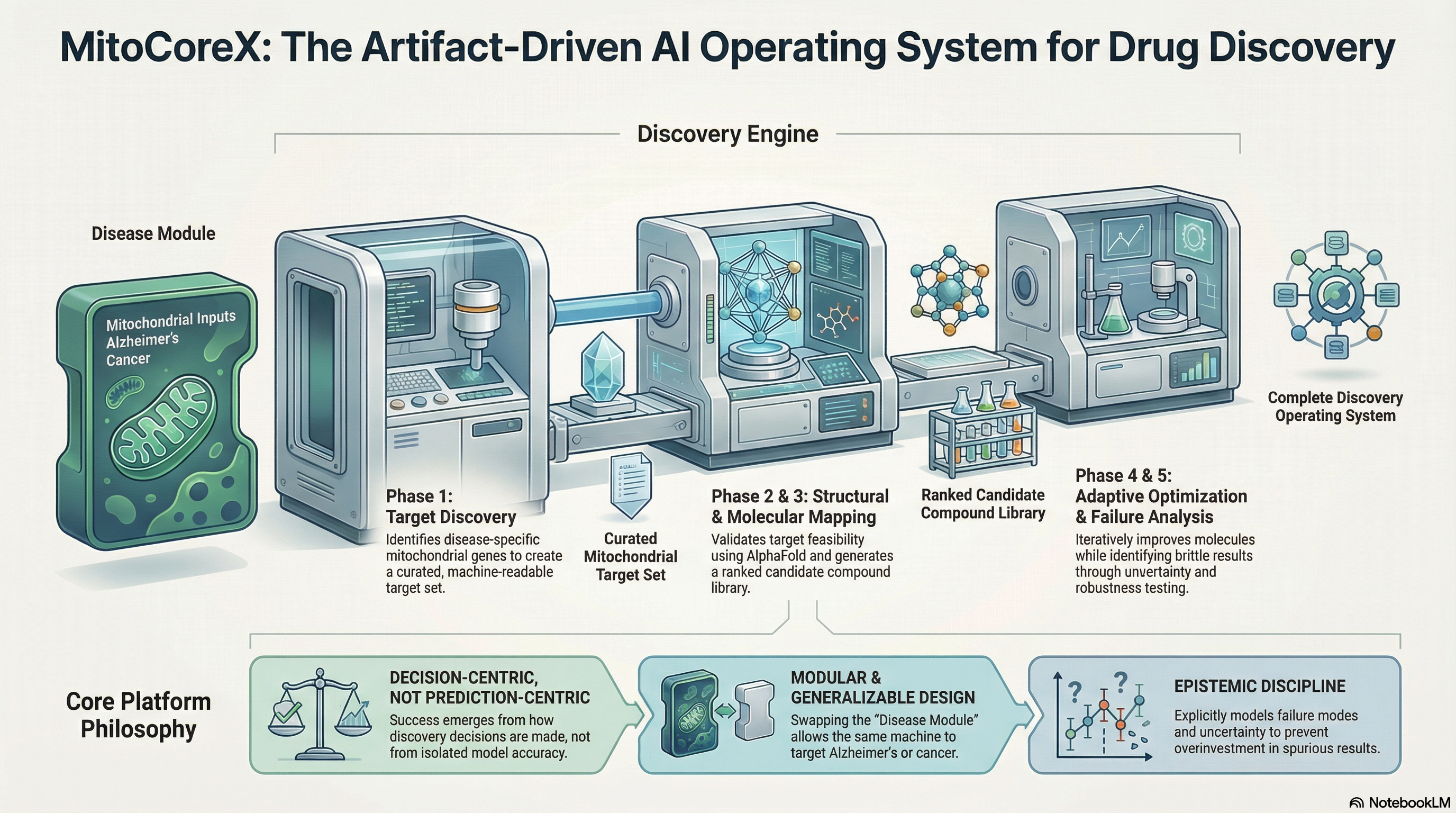
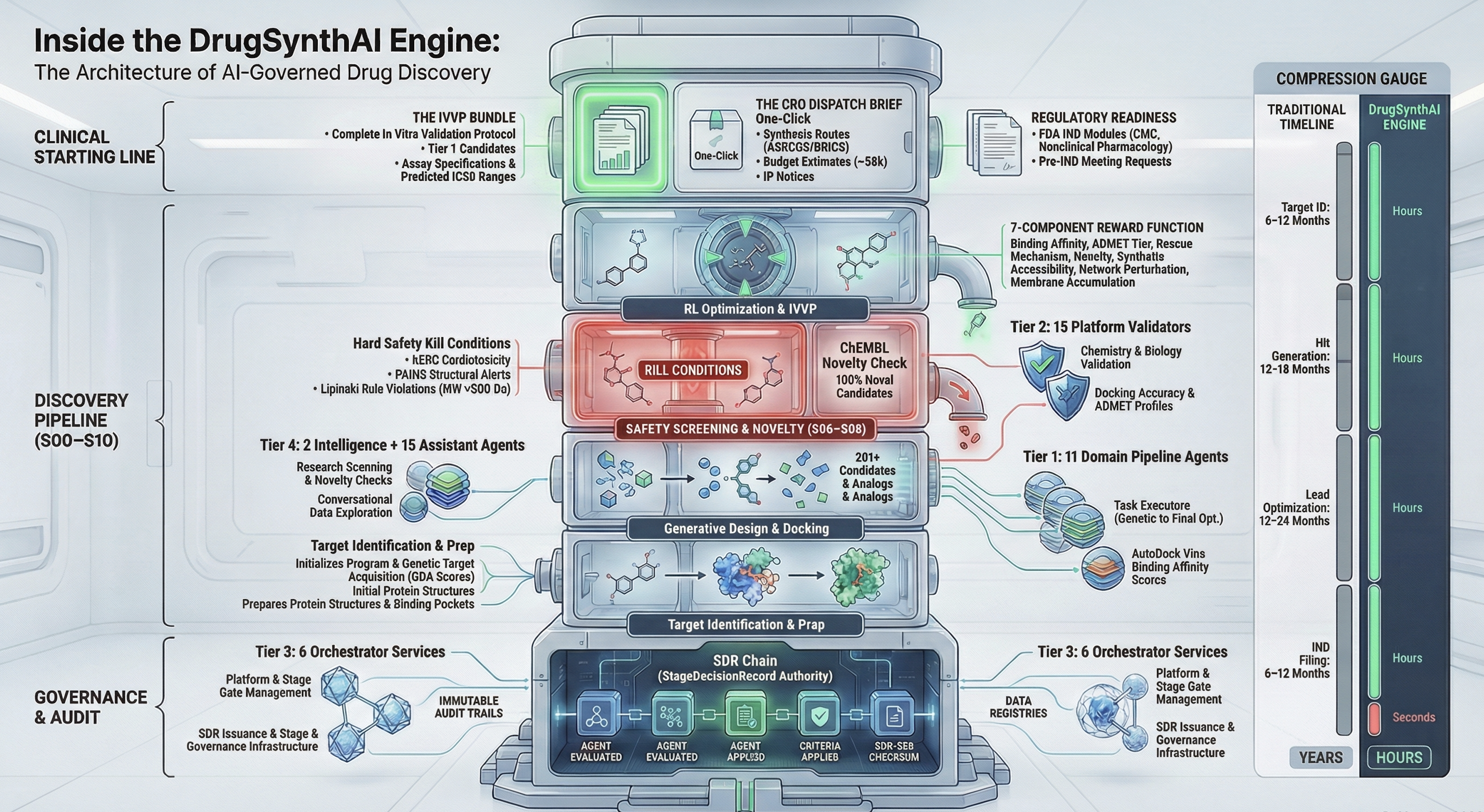
Other platforms do parts of this. None do all of it, and none do it with governance built in from the start. Click any row to see what was actually built.
CapabilityDrugSynthAIREINVENT 4Insilico
De novo molecule designYesYesYes▶
All three platforms generate novel molecules. DrugSynthAI uses RDKit BRICS fragment-based assembly with 90 curated fragments to produce 163 candidates per campaign, each with full 3D conformer generation. Molecules are true new chemical entities with computed SMILES, not database retrievals.
Real physics-based dockingYes (Vina 1.2.7)PredictedPredicted▶
DrugSynthAI runs AutoDock Vina 1.2.7 to compute actual binding free energies (ΔG in kcal/mol) against 22 docking-ready receptor structures with validated pocket geometries. Most competitors use machine learning models to predict binding scores. The difference matters for regulatory submissions: physics-based docking results are accepted by the FDA; ML predictions require additional experimental validation.
Governed pipeline with audit trailYes (AIDD-GOV)NoNo (proprietary)▶
Every stage transition produces a StageDecisionRecord (SDR) with SHA-256 checksums, agent attribution, and decision rationale. The audit trail is stored in append-only SQLite (WAL mode) and exportable as YAML. AIDD-GOV is published as an open standard (Apache 2.0, github.com/fxmedus/aidd-gov) with 10 JSON schemas and 3 conformance levels. No other platform publishes its governance logic as an auditable, adoptable standard.
Patient population mappingYes (54K patients)NoNo▶
The Patient Intelligence Layer maps 14 pathogenic genetic variants to 54,204 patients worldwide using four real-world evidence APIs (Orphanet, GARD/NIH, ClinVar, HPO). For each variant, the platform computes estimated patient counts, geographic distribution, and clinical presentation. This is not a static database: the platform queries live APIs at pipeline execution time and generates population estimates dynamically for any disease in its registry.
Regulatory document generationYes (Pre-IND, CMC, IND)NoNo▶
Five API endpoints generate FDA-format regulatory documents in seconds: a Pre-IND Type B meeting request letter with 9-section briefing document and 7 to 9 proposed questions; CMC modules (3.2.S drug substance + 3.2.P drug product) with specifications tables, analytical methods, and stability protocols; nonclinical summaries (Module 2.4 overview + 2.6 written summary) covering a 5-study toxicology program and 4-study PK program; and a complete IND package that assembles everything in one call. All structured to ICH Common Technical Document format.
CRO synthesis dispatchYesNoNo▶
When a compound graduates from the computational pipeline, the platform generates a contract research organization (CRO) dispatch brief: a structured document containing the retrosynthetic route (ASKCOS with BRICS fallback), starting material commercial availability, estimated synthesis difficulty, required analytical methods, and quality specifications. Available as JSON (machine-readable) or Markdown (human-readable). This bridges the gap between in silico design and physical synthesis, something no competitor automates.
Companion diagnostic designYesNoNo▶
For each genotype-to-compound match, the platform designs a 3-tier companion diagnostic panel: Tier 1 PCR (targeted single-gene, $250, 7-day turnaround), Tier 2 NGS panel (multi-gene, for complex variants), and Tier 3 WES/WGS (whole exome or genome, for novel mutations). The diagnostic specification includes primer targets, sensitivity estimates, and cost projections. This is precision medicine infrastructure: the right test to find the right patient for the right drug.
Delivery engineeringYes (7 strategies)NoPartial▶
Mitochondria are inside cells, behind two membranes. Getting a drug there is a delivery problem, not a chemistry problem. DrugSynthAI implements three delivery modules: TPP+ conjugation (triphenylphosphonium targeting, 850x mitochondrial accumulation), prodrug design (7 strategies including ester, amide, carbamate, phosphate, PEG, amino acid, and lipid prodrugs), and formulation specification (5 administration routes: IV, oral, intrathecal, topical, inhalation, each with CMC-grade excipient specifications). Each module generates structured output with estimated bioavailability improvements.
Molecular dynamics simulationYes (OpenMM)NoNo▶
Beyond static docking, the platform runs molecular dynamics simulations to evaluate binding stability over time: system preparation (PDBFixer + TIP3P water + AMBER14 force field), NVT/NPT equilibration, production MD with RMSD/RMSF analysis, and free energy perturbation (FEP) for ΔΔG ranking of lead candidates. This answers the question docking cannot: does the molecule stay bound, or does it fall out of the pocket?
Open governance standardYes (Apache 2.0)NoNo▶
AIDD-GOV is not just a feature of DrugSynthAI. It is an independent, open-source governance standard published for adoption by any AI drug discovery platform. The specification defines 10 JSON schemas covering StageDecisionRecords, constraint policies, convergence criteria, and audit event formats, with 3 conformance levels (Bronze, Silver, Gold). It is the first open standard for governing AI decisions in pharmaceutical research.
Self-improving governanceYes (GOV-FM)NoNo▶
GOV-FM (Governance Foundation Model) is a self-improving recursive governance scorer. After every pipeline campaign, it evaluates 5 dimensions (decision provenance, constraint integrity, optimization transparency, audit completeness, regulatory alignment), identifies governance gaps, generates actionable recommendations, and persists them to a governance memory file that loads at the start of the next campaign. With 207 training examples and growing, the platform gets more disciplined with every run, not less. This is governance that learns.
Real-world evidence integrationYes (4 APIs)NoNo▶
The platform queries four public real-world evidence APIs at runtime: Orphanet (rare disease classifications and epidemiology), GARD/NIH (genetic and rare disease information), ClinVar (NCBI clinical variant significance), and HPO (Human Phenotype Ontology for symptom mapping). This data feeds into patient population estimates, diagnostic panel design, and regulatory justification for rare disease designations (Orphan Drug, Fast Track, Breakthrough Therapy). No manual literature review needed.
Foundation/grant proposal generationYesNoNo▶
For rare disease programs, the platform generates structured foundation proposal sentences using patient population data, variant-to-compound mappings, and diagnostic panel specifications. These are formatted for patient advocacy foundations and NIH grant applications: prevalence data, unmet medical need justification, and proposed therapeutic approach, all backed by the platform's computational evidence and traceable to specific SDRs in the audit trail.
Patient-specific genomic intakeYes (VCF, RNA-seq)NoNo▶
Clinicians upload actual patient molecular data: VCF files, FoundationOne or Tempus clinical genomics reports, RNA-seq expression matrices, and methylation arrays. The Data Ingest Agent (DIA_001) parses these into a unified patient molecular profile, identifies actionable genetic variants, and maps them to the platform's 300-disease gene database automatically. No other drug discovery platform accepts patient-level molecular data as a campaign input.
Patient regulatory graph inferenceYes (GRN + perturbation)NoNo▶
The GRN Inference Agent (GRN_001) builds a patient-specific gene regulatory network from multi-omic data, weighted by enhancer-promoter interactions (Activity-by-Contact model) and transcription factor binding site predictions (JASPAR, HOCOMOCO). The Perturbation Agent (PRT_001) then simulates gene knockdowns, enhancer silencing, and pathway inhibition to identify highest-impact therapeutic targets with lowest off-target risk. This is not a textbook pathway: it is constructed from the patient's own expression data.
Multi-modal therapeutic generationYes (3 modalities)NoPartial▶
Within a single governed campaign, the platform generates three classes of therapeutic candidates: small molecules (163 per campaign, AutoDock Vina physics-based docking), peptides (50-fragment library, mitochondria-targeting sequences, cell-penetrating peptides, stapling strategies), and antibodies (CDR loop design, humanization scoring, developability profiling, immunogenicity prediction). All three modalities share the same governance trail, audit infrastructure, and kill condition enforcement. No other platform combines multi-modal generation under unified governance.
Patient therapeutic reportYes (PDF)NoNo▶
A structured report showing the patient's actionable variants, inferred regulatory dysfunction, top therapeutic candidates across all three modalities, confidence scores, safety flags, and recommended next steps. HIPAA-compliant: patient data is processed in-session and not stored after report generation. No PHI retained on platform servers. This closes the gap between "we know the genetic cause" and "here is a candidate molecule that addresses it."